 The standard KASP thermal cycling protocol has 10 cycles of touchdown PCR (annealing 61☌ to 55☌, decreasing 0.6☌ per cycle), then 26 cycles of standard 2-step PCR at the lower annealing temperature (55☌). The regions that were originally amplified during the highly specific early touchdown cycles will be further amplified and outcompete any non-specific amplification that may occur at the lower temperatures. The annealing temperature is gradually decreased to increase the efficiency of the reaction. The higher annealing temperatures in the early cycles of a touchdown ensure that only very specific base pairing will occur between the DNA and the primer, hence the first sequence to be amplified is most likely to be the sequence of interest. A touchdown PCR involves starting with a high annealing temperature and incrementally decreasing the annealing temperature each PCR cycle. The temperature used for the annealing stage determines the specificity of the reaction and hence the ability of the primers to anneal to the DNA template. Nucleic Acids Res. 1990 18(21):6409-6412.KASP™ chemistry utilises a two-step touchdown PCR method, with the elongation and annealing steps incorporated into a single step. Touchdown PCR uses a cycling program where the annealing temperature is gradually reduced (e.g. Optimization of the annealing temperature for DNA amplification in vitro. It is a method for increasing specificity of PCR reactions. Understand the importance of melting temperature in molecular biology applications In this type of PCR reaction, the initial or the annealing temperature is kept higher as compared to the standard temperature and this. Optimal annealing temperatures give the highest product yield of the correct amplicon. Conversely, when T a is too high reaction efficiency may be reduced because the likelihood of primer annealing is reduced significantly. This can lead to nonspecific PCR amplification and will consequently reduce the yield of the desired product. One consequence of having T a too low is that one or both primers will anneal to sequences other than the intended target, because internal single-base mismatches or partial annealing may be tolerated. To accomplish this, three significant changes were made 1) touchdown PCR cycling parameters were used to eliminate the need for optimizing PCR cycling conditions, 2) the high-fidelity, high-processivity Q5 DNA polymerase was used to improve full-length amplification quality, and 3) a reduced amount of primer in the final PCR amplification step. The optimal annealing temperature (T a Opt) for a given primer pair on a particular target can be calculated as follows: T a Opt = 0.3 x (T m of primer) + 0.7 x (T m of product) – 14.9, where T m of primer is the melting temperature of the less stable primer-template pair, and T m of product is the melting temperature of the PCR product. Generally, you should use an annealing temperature about 5☌ below the T m of your primers. In touchdown PCR, the PCR cycle is started with an annealing temperature slightly higher than the calculated or anticipated annealing temperature of the. The annealing temperature (T a) chosen for PCR relies directly on length and composition of the primers. Target Capture Probe Design & Ordering Tool.Library Concentration Conversion Calculator. 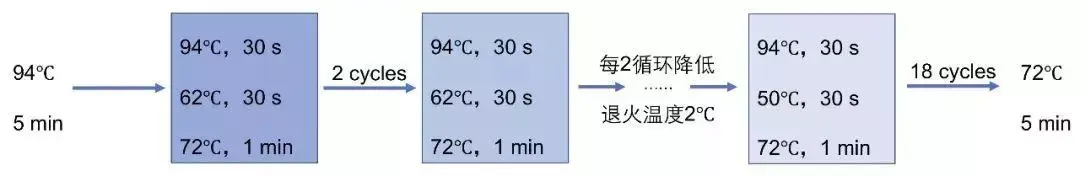
Alt-R Predesigned Cas9 crRNA Selection Tool.SYBR Green dye assay and PrimeTime probe assays.PCR Allele Competitive Extension (PACE) genotyping.Shotgun metagenomics for infectious diseases.Drug target identification via CRISPR screening. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
